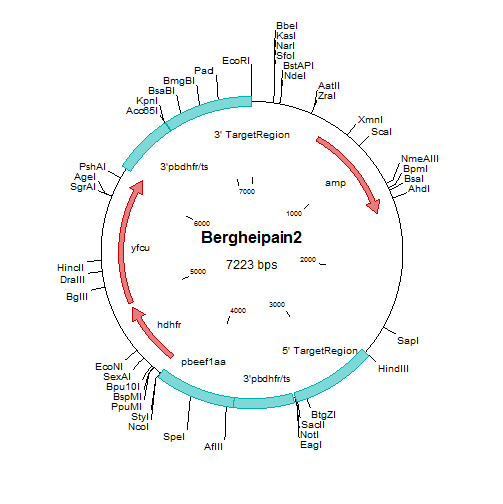
| Additional remarks phenotype | Mutant/mutation
The mutant lacks expression of berghepain-2 (BP2) and expresses the GFP-Luciferase fusion protein under control of the (schizont specific) ama-1 promoter.
Protein (function)
Berghepain-2 (BP2) is equivalent to the three P. falciparum digestive vacuole falcipains (papain-like cysteine endoproteases), FP-2 (FP2a, PF3D7_1115700; FP2b, PF3D7_1115300) and -3 (PF3D7_1115400). The gene encoding BP2 has a syntenic location to the three falcipain genes of P. falciparum and also the the single copy vivapain-2 of P. Vivax (PVX_086040).
Both P. falciparum and P. berghei have an additional papain-like cysteine protease encoded by a syntenic gene (falcipain 1, FP1, PF3D7_1458000 and berghepain 1, BP1, PBANKA_132170). Evidence has been presented that this protease is located (and active) in the (apical end) of merozoites and not in the digestive vacuole, suggesting that it has no role in hemoglobin digestion.
P. falciparum parasites catabolize more than half of the hemoglobin (Hb) in the RBC. The amino acids derived from Hb proteolysis are used for protein synthesis and energy metabolism. The proteolysis of Hb is accompanied by the release of free heme, which is cytotoxic for the parasite and is rapidly detoxified and converted into hemozoin (Hz). Therefore, both Hb degradation and heme detoxification are considered to be essential for P. falciparum survival. The digestion of Hb is a conserved and semi-ordered process, which principally occurs within the acidified digestive vacuole (DV). After the initial cleavage by by aspartic and papain-like cysteine endoproteases, Hb unfolds and becomes accessible to further proteolysis by downstream proteases. In the P. falciparum DV there are four aspartic proteases (plasmepsins I-IV) and two papain-like cysteine proteases (falcipains 2a,b and 3) capable of hydrolyzing native Hb. Gene disruption studies of hemoglobinases demonstrated that P. falciparum has developed redundant and overlapping Hb degradation pathways demonstrating the importance of Hb digestion for the parasite.
Evidence (from gene deletion studies) has been provided that FP2 is not essential for P. falciparum blood stage growth, whereas FP3 appears to be essential/
Phenotype
The phenotype analyses indicate that BP2 is not essential for blood stage growth.
The observations on Hz production in the single gene-deletion mutants Δpm4 (RMgm-808) and Δbp2 indicate that also in P. berghei aspartyl and cysteine endopeptidases overlap in their ability to hydrolyse Hb. Interestingly, the Δbp2 mutant has a normal growth rate and produces wt-levels of Hz, whereas Δpm4 parasites have a reduced growth and Hz production. These observations demonstrate that while PM4 is able to fully compensate for the function of BP2, BP2 can only partly compensate for the loss of PM4.
See also the 'double' gene deletion mutant RMgm-817 which lacks expression of both PM4 and BP2. Blood stages of this mutant show a strongly reduced hemoglobin digestion and hemozoin formation and parasites can grow and multiply in reticulocytes without the formation of hemozoin.
Additional information
An independent mutant (Pain2cl8, ∆bp2-a) was generated in the 1037m1f1cl1 reference P. berghei ANKA line in the group of Roberta Spaccapelo (University of Perugia, Italy) using a circular, double cross-over targeting plasmid (pLTgPain2). The 5' targeting region was amplified using primer pair R443/R444 (CCGGGCCCGCGGGGTTTCTATCTATATTTATTTCTGC / CCATCGATTTATGTTTCTATGTTAATTTTTTTTTGC). The 3' targeting region was amplified using primer pair R445/R446 (GGAATTCAAATAATATTATGTACCGATAGG / CGGGATCCTGGAATCGCCCTTTTATAATGC).
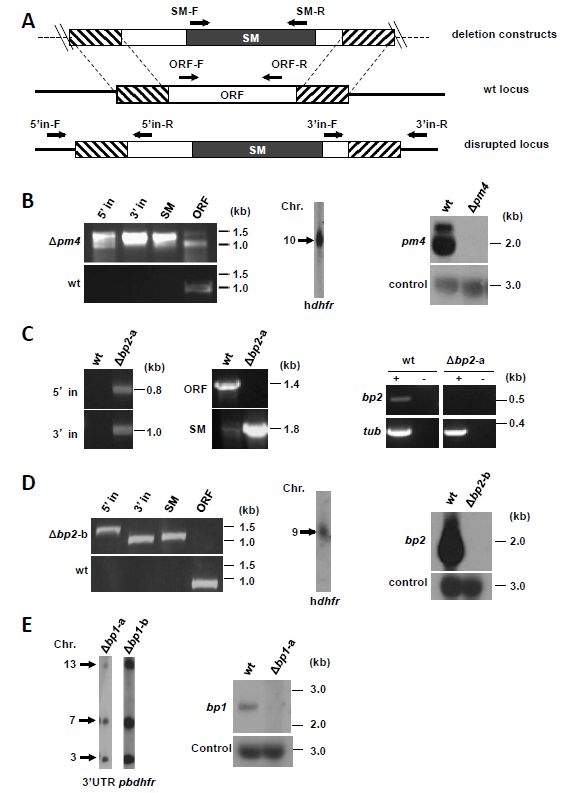
Figure: Generation of the P. berghei mutants ∆pm4 (RMgm-808), ∆bp1 (RMgm-816) and ∆bp2 (RMgm-809).
(primer sequences can either be found in Lin et al. (2015). J. Exp. Med. or obtained from the Leiden malaria Research Group).
(A) Schematic representation of gene-deletion constructs targeting the open reading frame (ORF) of genes expressing plasmepsin 4 (pm4), berghepain 2 (bp2) or berghepain 1 (bp1) by double cross-over homologous recombination, and wild-type (wt) gene loci before and after disruption. The constructs contain a drug-selectable marker cassette (SM; black box) and gene target regions (hatched boxes). Primer positions (arrows) for diagnostic PCRs are shown (see Table S4 for primer sequences and expected product sizes). (B) Diagnostic PCR (left) and Southern analysis of pulsed field gel-separated chromosomes (center) confirm correct disruption of pm4 in mutant ∆pm4-b. Northern analysis of blood-stage mRNA (right) confirms the absence of pm4 transcripts in the ∆pm4-b mutant. The following primers were used for diagnostic PCRs: 5' integration (5’ in): L5516/L4096; 3' integration: (3’ in) L1662/L5517; SM (hdfhr::yfcu): L4698/L4699; pm4 ORF: L5518/L5519. Separated chromosomes were hybridized using an hdhfr probe that recognizes the DNA-construct integrated into the pm4 locus on chromosome 10. Northern blot was hybridized using a PCR probe recognizing the pm4 ORF (primers L5518/L5519) and with an oligonucleotide probe L644R recognizing the large subunit rRNA (as loading control). (C) Diagnostic PCR (left) confirms the correct deletion of the bp2 gene in mutant ∆bp2-a. RT-PCR analysis of blood stage mRNA (right) shows the absence of bp2 transcription in ∆bp2-a blood-stages. The following primer pairs were used for diagnostic PCR analyses: 5’ in, RS835/RS32; 3’ in, RS110/RS836; SM (tgdhfr/ts), RS404/RS405; bp2 ORF, RS514/RS515. For RT-PCR the following primers were used: tub (tubulin), RS782/RS783 and bp2, RS515/RS516. (D) Diagnostic PCR (left) and Southern analysis of pulsed field gel-separated chromosomes (center) confirm the correct disruption of the bp2 gene in mutant ∆bp2-b. Northern blot analysis of blood stage mRNA (right) confirms the absence of bp2 transcripts in ∆bp2-b. The following primers were used for diagnostic PCRs: 5’ in, L5024/L3211; 3’ in, L5025/L1662; SM (hdhfr::yfcu), L4698/L4699; bp2 ORF, L5026/L5027. Separated chromosomes were hybridized using an hdhfr probe that recognizes the DNA-construct integrated into the bp2 locus on chromosome 9. Northern blot was hybridized using a PCR probe recognizing the bp2 ORF (primers L5026/L5027) and with an oligonucleotide probe L644R recognizing the large subunit rRNA (as loading control). (E) Southern analysis of pulsed field gel-separated chromosomes (left) confirms the correct disruption of bp1 in mutant ∆bp1. Northern analysis of blood-stage mRNA (right) confirms the absence of bp1 transcripts in mutant ∆bp1. Separated chromosomes were hybridized using an 3’UTR pbdhfr probe that recognizes the DNA-construct integrated into the bp1 locus on chromosome 13, the endogenous dhfr/ts on chromosome 7 and the GFP-luciferase reporter cassette in the 230p locus on chromosome 3. Northern blot was hybridized using a PCR probe recognizing the bp1 ORF (primers L7422/L7423) and with an oligonucleotide probe L644R recognizing the large subunit rRNA (as loading control). See Table S4 for primers used for generation of probes.
Other mutants
RMgm-816: a mutant lacking expression of BP1
RMgm-817: a 'double' gene deletion mutant lacking expression of both PM4 (Plasmepsin 4) and BP2 (berghepain-2)
See the link for other 'plasmepsin 4' mutants
|
 *RMgm-809
*RMgm-809 Disrupted: Mutant parasite with a disrupted gene
Disrupted: Mutant parasite with a disrupted gene




 Transgene: Mutant parasite expressing a transgene
Transgene: Mutant parasite expressing a transgene