Back to search resultsSummaryRMgm-608
|
||||||||
 *RMgm-608
*RMgm-608| Successful modification | The parasite was generated by the genetic modification |
| The mutant contains the following genetic modification(s) | Gene mutation |
| Reference (PubMed-PMID number) |
Reference 1 (PMID number) : 21262960 |
| MR4 number | |
| top of page | |
| Parent parasite used to introduce the genetic modification | |
| Rodent Malaria Parasite | P. berghei |
| Parent strain/line | P. berghei ANKA |
| Name parent line/clone | Not applicable |
| Other information parent line | |
| top of page | |
| The mutant parasite was generated by | |
| Name PI/Researcher | A. Coppi; P. Sinnis |
| Name Group/Department | Department of Medical Parasitology |
| Name Institute | New York University School of Medicine |
| City | New York |
| Country | USA |
| top of page | |
| Name of the mutant parasite | |
| RMgm number | RMgm-608 |
| Principal name | ∆RI |
| Alternative name | |
| Standardized name | |
| Is the mutant parasite cloned after genetic modification | Yes |
| top of page | |
| Phenotype | |
| Asexual blood stage | Not different from wild type |
| Gametocyte/Gamete | Not different from wild type |
| Fertilization and ookinete | Not different from wild type |
| Oocyst | Not different from wild type |
| Sporozoite | Normal numbers of salivary gland sporozoites are produced. Mutant sporozoites showed impaired ability (fivefold reduction) to invade hepatocytes in vitro. Motility and cell traversal of mutant sporozoites were normal. |
| Liver stage | Mutant sporozoites showed impaired ability (fivefold reduction) to invade hepatocytes in vitro. Liver stages (EEF's) that were formed in vitro showed normal morphology. Mutant sporozoites, intravenously injected into mice, showed a 15-fold reduction in infectivity compared to wild type sporozoites. A comparable reduction of infectivity was observed after intradermal inoculation of sporozoites. |
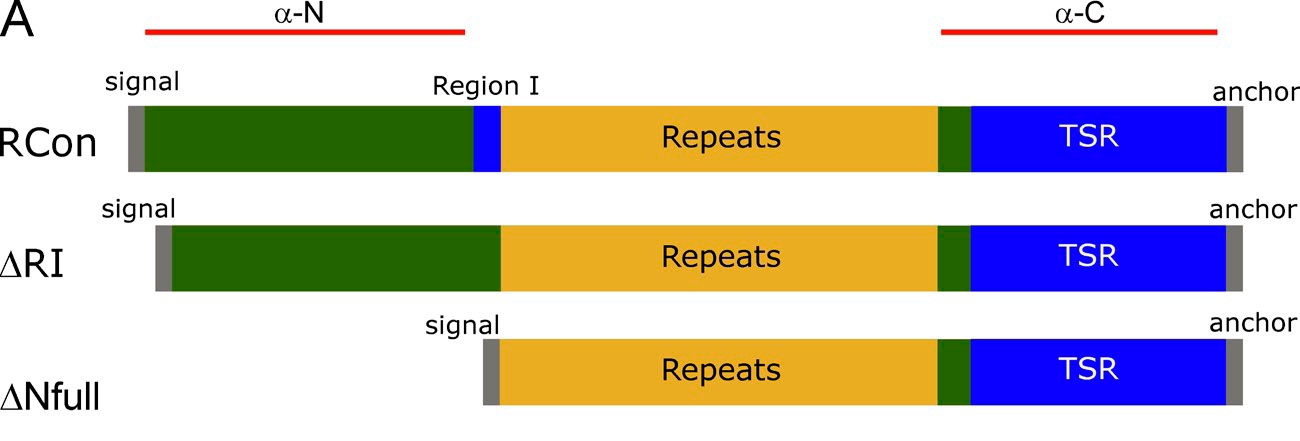
| Additional remarks phenotype | Mutant/mutation Overall, the phenotypic analyses suggest that the CSP cleavage site lies within the highly conserved 5-aa sequence called region I and demonstrate that region I is required for rapid and complete processing of CSP. The phenotype of ΔRI sporozoites establishes the link between CSP cleavage and hepatocyte invasion and demonstrates that efficient cleavage requires region I. Moreover, evidence is presented that proteolytic processing of CSP is required for TSR exposure in the mammalian host, i.e., during hepatocyte invasion, but not for its exposure during sporozoite development in the mosquito host. The TSR domain is a known cell-adhesive motif C-terminal to the repeats termed the type I thrombospondin repeat (TSR). See also mutant ∆Nfull (RMgm-609) in which the wild type csp gene is replaced with a mutated csp gene, lacking the N-terminal region (including Region I) excluding the signal sequence (see figure below)
Figure: Mutated csp genes from the different P. berghei mutant parasites as described in Coppi et al., 2011, J. Exp. Med. 208, 341-56. (RCon: wild type csp; ∆Nfull: see RMgm-609) |
 Mutated: Mutant parasite with a mutated gene
Mutated: Mutant parasite with a mutated gene| top of page | |||||||||||||||||||||||||||
| Details of the target gene | |||||||||||||||||||||||||||
| Gene Model of Rodent Parasite | PBANKA_0403200 | ||||||||||||||||||||||||||
| Gene Model P. falciparum ortholog | PF3D7_0304600 | ||||||||||||||||||||||||||
| Gene product | circumsporozoite (CS) protein | ||||||||||||||||||||||||||
| Gene product: Alternative name | CS, CSP | ||||||||||||||||||||||||||
| top of page | |||||||||||||||||||||||||||
| Details of the genetic modification | |||||||||||||||||||||||||||
| Short description of the mutation | deletion of region 1 (amino acids KLKQP) | ||||||||||||||||||||||||||
| Inducable system used | No | ||||||||||||||||||||||||||
| Short description of the conditional mutagenesis | Not available | ||||||||||||||||||||||||||
| Additional remarks inducable system | |||||||||||||||||||||||||||
| Type of plasmid/construct | Plasmid double cross-over | ||||||||||||||||||||||||||
| PlasmoGEM (Sanger) construct/vector used | No | ||||||||||||||||||||||||||
| Modified PlasmoGEM construct/vector used | No | ||||||||||||||||||||||||||
| Plasmid/construct map | |||||||||||||||||||||||||||
| Plasmid/construct sequence | |||||||||||||||||||||||||||
| Restriction sites to linearize plasmid | |||||||||||||||||||||||||||
| Selectable marker used to select the mutant parasite | hdhfr | ||||||||||||||||||||||||||
| Promoter of the selectable marker | pbdhfr | ||||||||||||||||||||||||||
| Selection (positive) procedure | pyrimethamine | ||||||||||||||||||||||||||
| Selection (negative) procedure | No | ||||||||||||||||||||||||||
| Additional remarks genetic modification | A mutant CSP gene was generated using a PCR-based approach. Two gene fragments flanking the region to be deleted and including engineered or endogenous restriction sites were amplified and cut to yield fragments that when ligated make a CSP mutant containing the desired deletion. For the ∆RI CSP, the amino acids KLKQP were deleted using the following strategy: a 28-bp 5’ fragment from the CSP open reading frame was amplified by PCR using forward primer P1 (5’-GTATCACGTGCTTAACTCTAAG-3’; existing PmlI site underlined) and P2 (5’-GCAATATTATTACGCTCTATTTTTTCG-3’; introduced SspI site underlined). Next, a 776-bp 3’ CSP fragment was amplified using forward primer P3 (5’- AGCGTAATAATAAATTGAAACAAAGGCCTCCACCACCAAACCC- 3’; introduced StuI site underlined) and reverse primer P4 (5’- TTATTTAATTAAAGAATACTAATAC-3’; existing PacI site underlined). Both PCR products were gel purified, digested SspI and StuI (resp.) to remove region 1, and ligated. The correct PmlI-PacI ligation product was determined by pcr and subsequently sub-cloned, sequenced and cloned into the pCSComp transfection plasmid. To generate the final pCSRep targeting construct that would replace the endogenous CSP locus, a CSP 5’UTR targeting region was cloned upstream of the selection cassette in pCSComp. The 5’UTR targeting region was obtained from p9.5∆E by PCR using p9.5∆E specific primers (see article) | ||||||||||||||||||||||||||
| Additional remarks selection procedure | |||||||||||||||||||||||||||
| |||||||||||||||||||||||||||
| top of page | |||||||||||||||||||||||||||